 References
References
Tutorial level: Introductory
Prior knowledge required: Suitable for biochemists with an understanding of biochemical networks, and enzyme kinetics, as well as for computer scientists and engineers familiar with basic modelling approaches.
Quantitative models of biochemical networks (signal transduction cascades, metabolic pathways, gene regulatory circuits) are a central component of modern systems biology. Building and managing these complex models is a major challenge that can benefit from the application of formal methods adopted from theoretical computing science. In this tutorial we provide a general introduction to the field of formal modelling, which emphasizes the intuitive biochemical basis of the modelling process, but is also accessible for an audience with a background in computing science and/or model engineering.
When building an ODE model, model complexity can rapidly increase to a level that is difficult to manipulate. Computational tools that allow the modular construction and visualization of ODE models would be very helpful. In addition, one usually faces a number of non-trivial design choices during model building. Even for a relatively simple model there may be many ways to describe its dynamic behaviour.
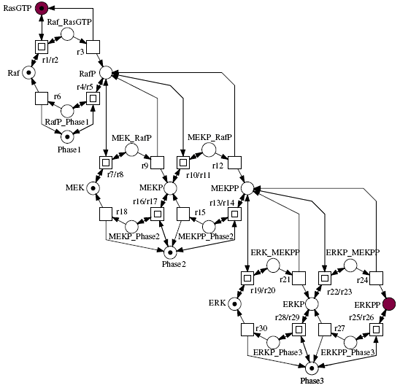
In this tutorial we show how signal transduction cascades can be modelled in a modular fashion, using both a qualitative approach – Qualitative Petri nets, and quantitative approaches -- Continuous Petri Nets and Ordinary Differential Equations. We review the major elementary building blocks of a cellular signalling model, discuss which critical design decisions have to be made during model building, and present a number of novel computational tools that can help to explore alternative modular models in an easy and intuitive manner. These tools, which are based on Petri net theory, offer convenient ways of composing hierarchical ODE models, and permit a qualitative analysis of their behaviour.
We will illustrate our approach using a generic model of a signalling cascade, and then relate this to existing models of the MAPK pathway, e.g. Levchenko, Brown and Schoerbel. We will show how the dynamic behaviour of such a pathway is related to its modular structure, and explore the use of stochastic versus deterministic techniques to generate behaviour traces. We will also introduce the use of temporal logic model checking of the pathway to characterise behavioural properties, using software recently developed at the University of Glasgow.
The ultimate aim is to introduce a general approach that provides the foundations for a structured formal engineering of large-scale models of biochemical networks.
We will illustrate the construction of the models in Ordinary Differential Equations using the BioNessie workbench from the University of Glasgow. This SBML compatible tool permits model construction of the models with a biochemistry-oriented interface, parameter scanning, fitting, and model checking using linear temporal logic. We will also illustrate the use of MatLab and the SimBio toolbox for the construction of models, which permits the use of the powerful programming and analytical facilities of MatLab. Petri net approaches will be done using the Snoopy and Charlie tools, both from the Brandenburg University of Technology Cottbus. Snoopy supports qualitative as well as quantitative Petri nets, including continuous and stochastic Petri nets. Snoopy's export feature permits interfacing to various analysis tools devoted to standard Petri net theory, as well as a variety of model checkers. There is also an export allowing access to other tools such as BioNessie, MatLab and the Glasgow model checker permitting more detailed evaluations of continuous and stochastic Petri nets in addition to the standard algorithms of ODE solvers provided by Snoopy.
 References
References
Software
Models
Tutorial URL: http://people.brunel.ac.uk/~csstdrg/workshops/APBC2009/
News
Jobs
The
School of Information Science, Computing and Mathematics at
Brunel University
in London, UK
is strenthening its activities in Bioinformatics and Systems Biology, as well
as establishing new activities in Synthetic Biology.
There are several open positions for talented and able
postdoctoral researchers and PhD students in
the area of Computational Systems and Synthetic Biology at Brunel.
For more details, see
David Gilbert's website.
The Groningen Bioinformatics Centre
at the University of Groningen, The
Netherlands, is looking for creative bioinformaticians with an interest in
Systems Biology, Metabolomics, Proteomics, Quantitative Genetics, Network
Reconstruction, Dynamic Modelling... to expand its young and successful
team.
Several PhD and Postdoc positions are available immediately.
For more information on our recent work, have a look at Rainer
Breitling's website. There
is also a partial list of GBiC
vacancies at the GBiC homepage.
If you are interested in a career in Groningen, please try to talk to
Rainer.
The International Max Planck Research School for Analysis, Design and Opt imization in Chemical and Biochemical Process Engineering in Magdeburg/Germany (IMPRS), offers young researchers a stimulating and challenging atmosphere for education and research in a highly international and interdisciplinary environment. For open positions (PhD students and postdoctoral researchers) see Monika Heiner's website and of her IMPRS cooperation partner Wolfgang Marwan.
Sponsors
The work presented in this tutorial is partially funded under the 6th Framework Programme by the European
Commission
(Integrated biomedical information for better health),
within the context of the SIMAP project.


